SUIT is a Matlab toolbox dedicated to the analysis of imaging data of the human cerebellum. The toolbox contains a high-resolution atlas template of the human cerebellum and brainstem, based on the anatomy of 20 young healthy individuals. The toolbox allows you to...
- ...automatically isolate cerebellar structures from the cerebral cortex based on an anatomical image.
- ...achieve accurate anatomical normalisation of cerebellar structures into atlas space using the Dartel algorithm.
- ...display the functional data on a surface-based flatmap representation.
- ...normalize focal cerebellar lesions for lesion-symptom mapping.
- ...use Voxel-based morphometry (VBM) to determine patterns of cerebellar degeneration or growth.
- ...use a probabilistic atlas of cerebellar anatomy to assign locations to different cerebellar lobules and deep cerebellar nuclei.
- ...use a collection of functional atlases based on task-based data and task-free connectivity to define regions of interest.
- ...improve normalization of the deep cerebellar nuclei using an ROI-driven normalization.
Please cite the use of this template as:
- Diedrichsen, J. (2006). A spatially unbiased atlas template of the human cerebellum. Neuroimage, 33, 1, p. 127-138.

- Diedrichsen, J., Balsters, J. H., Flavell, J., Cussans, E., & Ramnani, N. (2009). A probabilistic atlas of the human cerebellum. Neuroimage.

-
Diedrichsen, J., Maderwald, S., Kuper, M., Thurling, M., Rabe, K., Gizewski, E. R., et al. (2011). Imaging the deep cerebellar nuclei: A probabilistic atlas and normalization procedure. Neuroimage.

-
Diedrichsen, J. & Zotow, E. (2015). Surface-based display of volume-averaged
cerebellar data. PLoS One, 7, e0133402.

The toolbox has been developed by: J. Diedrichsen & C. Hernandez-Castillo, with help from M. King, G. Prichard, N. Lally, T.Wiestler, J. Schlerf, and many others.
Why a specialized template?
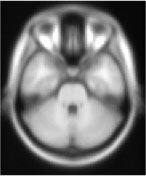
ICBM 152 Template
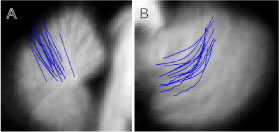
Superpostion of fissures after alignment to MNI template
The most commonly used atlas template for brain imaging is the ICBM152 template, defining the so-called MNI space. The template was generated by averaging 152 individual brains after affine normalization (correcting for size, translation and rotation). As it can be seen on the left, the template provides very little contrast for cerebellar structures. Normalization (affine or nonlinear) to this template leads to a large spatial spread of individual fissures. On the right is depicted the superposition of 16 individual primary fissures (A) and intra-biventer fissures (B). The distance between identical fissures in two different individuals is 4 mm on average.
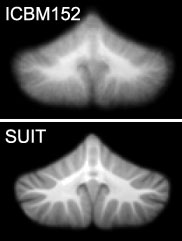
SUIT template and ICBM152 compared
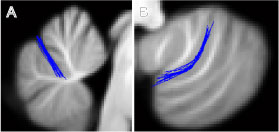
Superposition of fissures after alignment to SUIT template
We therefore developed a new template of the cerebellum and brainstem, the spatially unbiased infra-tentorial template (SUIT). This template is:
- based on the average anatomy of 20 young, healthy individuals, aged from 22 to 45
- preserves the anatomical detail of the cerebellum to a much higher degree than the MNI whole-brain template (see right).
- improves the overlap of the primary fissure (A, below) and the intra-biventer fissure (B, below), dramatically reducing the spatial variance
- improves the overlap of the deep cerebellar nuclei, although these structures are not visible on a T1 scan
Spatial unbiasedness, and the relationship to MNI space
The SUIT template was developed to be spatially unbiased in respect to the affine alignment to MNI space. This means that when normalizing the same brain to the MNI whole-brain template and to the SUIT template, the same structure should end up on average at the same coordinate in atlas space. For each individual there will be of course differences between the two normalizations (otherwise what would be the point?). These differences can be as big as 1 cm, and are on average across the image ~5mm. Furthermore, small systematic differences between different MNI normalisation methods (SPM Segmentation, SUIT, FSL's FNIRT) still remain. Therefore for the most accurate results, it is important that your data and your atlas template are normalised with the same algorithm. For the surface-based representation and the probabilistic atlas, we have therefore released separate versions for different atlas spaces.
Surface-based representation of the human cerebellum
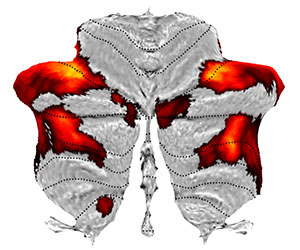
Flatmap representation
The SUIT toolbox also includes a surface-based representation of the human cerebellum (Diedrichsen & Zotow, 2015). This allows you to display volume-averaged cerebellar data on a group flatmap - providing an intuitive and efficient way to communicate cerebellar activation results. You can also explore common functional atlases on our interactive Atlas Viewer.